Algorithms¶
pyrecsys provides, out of the box, some basic algorithms based on matrix factorization.
SVD¶
pyrecsys makes use of SVD in order to decompose the input data (a matrix). Once the matrix is reduced into a lower dimensional space, pyrecsys can provide predictions, recommendations and similarity among the “elements” (being either users or items -it’s just a matter of how you load the matrix data-).
Loading data¶
How to load a dataset (e.g. Movielens 10M)?
from recsys.algorithm.factorize import SVD
filename = './data/movielens/ratings.dat'
svd = SVD()
svd.load_data(filename=filename, sep='::', format={'col':0, 'row':1, 'value':2, 'ids': int})
# About format parameter:
# 'row': 1 -> Rows in matrix come from second column in ratings.dat file
# 'col': 0 -> Cols in matrix come from first column in ratings.dat file
# 'value': 2 -> Values (Mij) in matrix come from third column in ratings.dat file
# 'ids': int -> Ids (row and col ids) are integers (not strings)
Split a dataset (train and test):
from recsys.datamodel.data import Data
from recsys.algorithm.factorize import SVD
filename = './data/movielens/ratings.dat'
data = Data()
format = {'col':0, 'row':1, 'value':2, 'ids': int}
data.load(filename, sep='::', format=format)
train, test = data.split_train_test(percent=80) # 80% train, 20% test
svd = SVD()
svd.set_data(train)
Loading a file, executing external SVDLIBC program, and creating an SVD instance svd:
from recsys.utils.svdlibc import SVDLIBC
svdlibc = SVDLIBC('./data/movielens/ratings.dat')
svdlibc.to_sparse_matrix(sep='::', format={'col':0, 'row':1, 'value':2, 'ids': int}) # Convert to sparse matrix format [http://tedlab.mit.edu/~dr/SVDLIBC/SVD_F_ST.html]
svdlibc.compute(k=100)
svd = svdlibc.export()
Computing¶
>>> K=100
>>> svd.compute(k=K, min_values=10, pre_normalize=None, mean_center=True, post_normalize=True, savefile=None)
Parameters:
min_values: remove those rows or columns (from the input matrix) that has less than ‘min_values’ non-zeros
pre_normalize: normalize input matrix. Possible values are tfidf, rows, cols, all.
tfidf: By default, treats the matrix as terms-by-documents. It’s important, then, how the data is loaded. Use the format param in svd.load_data() to determine the order of the fields of the input file.
rows: Rescales the rows of the input matrix so that they all have unit Euclidean magnitude
cols: Rescales the columns of the input matrix so that they all have unit Euclidean magnitude
all: Rescales the rows and columns of the input matrix, by dividing both the rows and the columns by the square root of their Euclidean norm
mean_center: centering the input matrix (aka mean substraction)
post_normalize: Normalize every row of
to be a unit vector. Thus, row similarity (using cosine distance) returns
savefile: Output file to store SVD transformation (
vectors)
Predictions¶
To predict a rating, 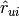 , SVD class reconstructs the original matrix,
, SVD class reconstructs the original matrix, 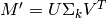
Then,
>>> svd.predict(ITEMID, USERID, MIN_RATING=0.0, MAX_RATING=5.0)
equals to:
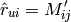
Here are the RMSE and MAE values for the Movielens 10M dataset (Train: 8,000,043 ratings, and Test: 2,000,011), using 5-fold cross validation, and different K values or factors (10, 20, 50, and 100) for SVD:
| K | 10 | 20 | 50 | 100 |
| RMSE | 0.87224 | 0.86774 | 0.86557 | 0.86628 |
| MAE | 0.67114 | 0.66719 | 0.66484 | 0.66513 |
Recommendations¶
Recommendations (i.e. unknown values in 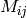 ) are also derived from
) are also derived from 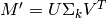 . In this case,
. In this case,
>>> svd.recommend(USERID, n=10, only_unknowns=True, is_row=False)
returns the higher values (e.g. items that the user has not rated) of 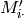
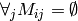 , whilst
, whilst
>>> svd.recommend(USERID, n=10, only_unknowns=False, is_row=False)
returns the best items for the user
Neighbourhood SVD¶
Classic Neighbourhood algorithm uses the ratings of the similar users (or items) to predict the values of the input matrix M.
from recsys.algorithm.factorize import SVDNeighbourhood
svd = SVDNeighbourhood()
svd.load_data(filename=sys.argv[1], sep='::', format={'col':0, 'row':1, 'value':2, 'ids': int})
K=100
svd.compute(k=K, min_values=5, pre_normalize=None, mean_center=True, post_normalize=True)
Predictions¶
The only difference with plain SVD is the way how it computes the predictions 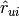
>>> svd.predict(ITEMID, USERID, weighted=True, MIN_VALUE=0.0, MAX_VALUE=5.0)
To compute the prediction, it uses this equation (u=USERID, i=ITEMID):
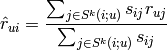
where
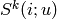 denotes the set of
denotes the set of 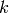 items rated by
items rated by  , which are most similar to
, which are most similar to 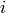 .
.
- To compute the
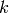 items most similar to
items most similar to 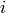 , it uses the svd.similar(i) method. Then it gets those items that user
, it uses the svd.similar(i) method. Then it gets those items that user  has already rated
has already rated
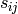 is the similarity between
is the similarity between 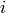 and
and 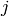 , computed using svd.similarity(i, j)
, computed using svd.similarity(i, j)
Comparison¶
For those who love RMSE, MAE and the like, here are some numbers comparing both SVD approaches. The evaluation uses the Movielens 1M ratings dataset, splitting the train/test dataset with ~80%-20%.
Note
Computing svd k=100, min_values=5, pre_normalize=None, mean_center=True, post_normalize=True
Warning
Because of min_values=5, some rows (movies) or columns (users) in the input matrix are removed. In fact, those movies that had less than 5 users who rated it, and those users that rated less than 5 movies are removed.
Results¶
Movielens 1M dataset (number of ratings in the Test dataset: 209,908):
| SVD | SVD Neigh. | |
| RMSE | 0.91811 | 0.875496 |
| MAE | 0.71703 | 0.684173 |
![[-1.0 .. 1.0]](_images/math/24323325974c3d0f493c95a63f6818fb012d7b0c.png)